Nearly 12,000 people per year die of an antibiotic-resistant infection called MRSA only in America. One of the main problems that mankind is dealing nowadays is the rapid growth of bacterial resistance. Bacteria are rapidly evolving resistance to existing drugs, making the need for newer, different and probably stronger antibiotics.
Now, the researchers from the Duke University think they’ve solved a piece of the puzzle of breaking bacterial resistance. They hope it could lead to the development of new broad-spectrum antibiotics.
The study published on Dec. 26 in Nature Structural and Molecular Biology, has provided the first close-up glimpse of a protein, called MurJ, which is crucial for building the bacterial cell wall and protecting it from outside attack.
Researchers explain that bacteria have an additional layer of protection, called the cell wall, that animal cells don’t have. This tough wall acts like the powerful armor for bacteria, making them almost indestructible.
But, there could be a key for breaking that protective bacterial wall and bacterial resistance generally.
“Until now, MurJ’s mechanisms have been somewhat of a ‘black box’ in the bacterial cell wall synthesis because of technical difficulties studying the protein,” said senior author Seok-Yong Lee, Ph.D., associate professor of biochemistry at Duke University School of Medicine.
“Our study could provide insight into the development of broad -spectrum antibiotics because nearly every type of bacteria needs this protein’s action.”
A bacterium’s cell wall is composed of a rigid mesh-like material called peptidoglycan. The biogenesis of this material is a key antibiotic target.
Researchers have noticed that certain molecules that manufacture peptidoglycan are formed inside the cell, but they need to be transported across the cell membrane to build the protective cell wall. MurJ protein is actually responsible for flipping these wall building blocks across the membrane.
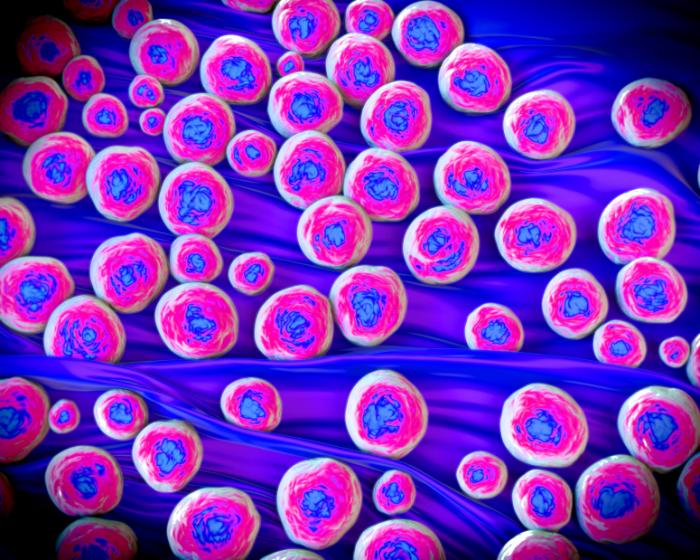
MRSA, full name methicillin-resistant staphylococcus aureus, is a form of bacterial infection that is resistant to numerous antibiotics including methicillin, amoxicillin, penicillin and oxacillin. Source: MedicalNewsToday
Without MurJ the forming of bacterial protection layer wouldn’t have been possible. The bacterium would fall apart.
But bacterial membrane proteins are complex and difficult to work with, but the team hopes it will understand the mechanism soon. So for now, understanding of how this process works remains limited due to a lack of structural information
“Getting the structure of MurJ linked to its substrate will be key. It will really help us understand how this transporter works and how to develop an inhibitor targeting this transporter,” Lee said.
Lee’s group is continuing structure and function studies of other key players in bacterial cell wall biosynthesis as well. Last year, they published the structure of another important enzyme, MraY, bound to the antibacterial muraymycin, ScienceDaily reports. The research was supported by Duke University startup funds.